|
IDs that can be recognized by BioVenn as belonging to a certain database, are linked to that database. In this article, we present a web application named BioVenn which can generate circular, area-proportional Venn diagrams just by entering two or three lists of biological IDs. There is currently no web application available that can generate circular, area-proportional Venn diagrams connected to a wide range of biological databases, and can map different kinds of IDs to genes. There is also the Google Chart API, which can generate circular, area-proportional Venn Diagrams, but can only have three numbers as input, and cannot do any calculations to obtain these three numbers. Drawback of these programs is that they need to be downloaded and run locally, limiting their use by a wide community. 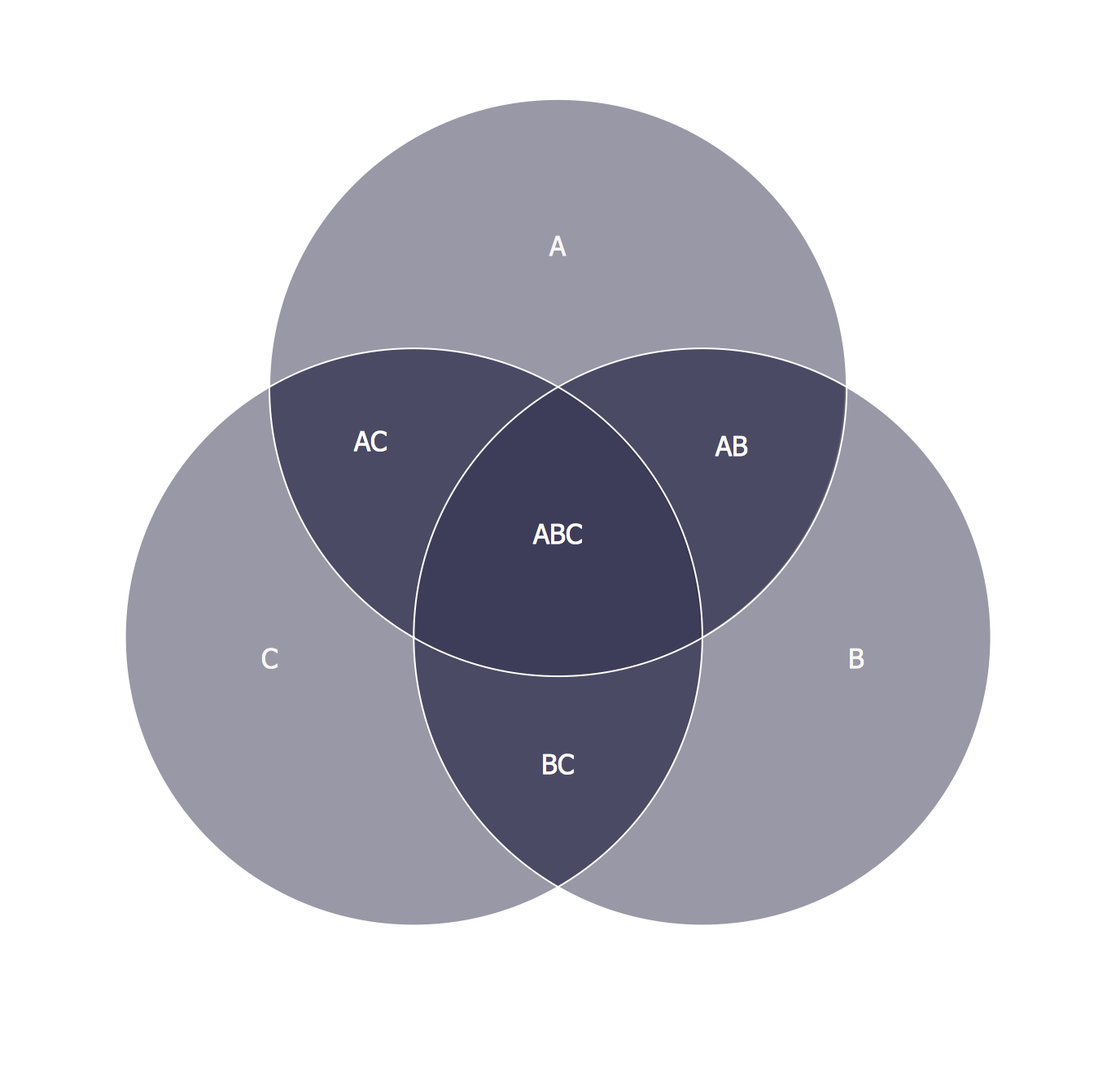
There are some computer programs available that generate area-proportional Venn Diagrams, either as rectangles or as polygons. However, these applications generate diagrams with circles of equal size. Venn diagrams have been used recently to visualize gene lists. This is called an area-proportional Venn diagram. In such a diagram, the size of the circle can be used to represent the size of the corresponding data set. A large number of different types of Venn diagrams exist, the most popular being the three-circle Venn diagram, used to visualize the overlap between three data sets. One of the most popular methods to visualize the overlap and differences between data sets is the Venn diagram, named by its inventor John Venn. This enables researchers to quickly observe similarities and differences between the data sets they are analyzing. Often, it is useful to see the overlap between these lists. sets of genes regulated under different treatments. In many genomics projects and other projects handling large amounts of biological data, various lists containing biological identifiers are produced, corresponding to e.g. Its implementation on the World Wide Web makes it available for use on any computer with internet connection, independent of operating system and without the need to install programs locally. It supports a wide range of identifiers from the most used biological databases currently available. ConclusionīioVenn is an easy-to-use web application to generate area-proportional Venn diagrams from lists of biological identifiers. Finally, BioVenn can map Affymetrix and EntrezGene identifiers to Ensembl genes. If an identifier is recognized as belonging to one of the supported biological databases, the output is linked to that database. Besides the Venn diagram, BioVenn outputs lists of identifiers for each of the resulting subsets. The latter option is useful for batch queries. The output Venn diagram can be shown as an SVG or PNG image embedded in the web application, or as a standalone SVG or PNG image. The position of the text can be adjusted by 'drag-and-drop' principle. Parameters like colors and text size can be adjusted easily through the web interface. 
The user only needs to input these lists of identifiers in the textboxes and push the submit button. We designed a web application named BioVenn to summarize the overlap between two or three lists of identifiers, using area-proportional Venn diagrams. Currently there are no programs available that can create area-proportional Venn diagrams connected to a wide range of biological databases. the sizes of the circles and the overlaps correspond to the sizes of the data sets. Venn diagrams are especially useful when they are 'area-proportional' i.e.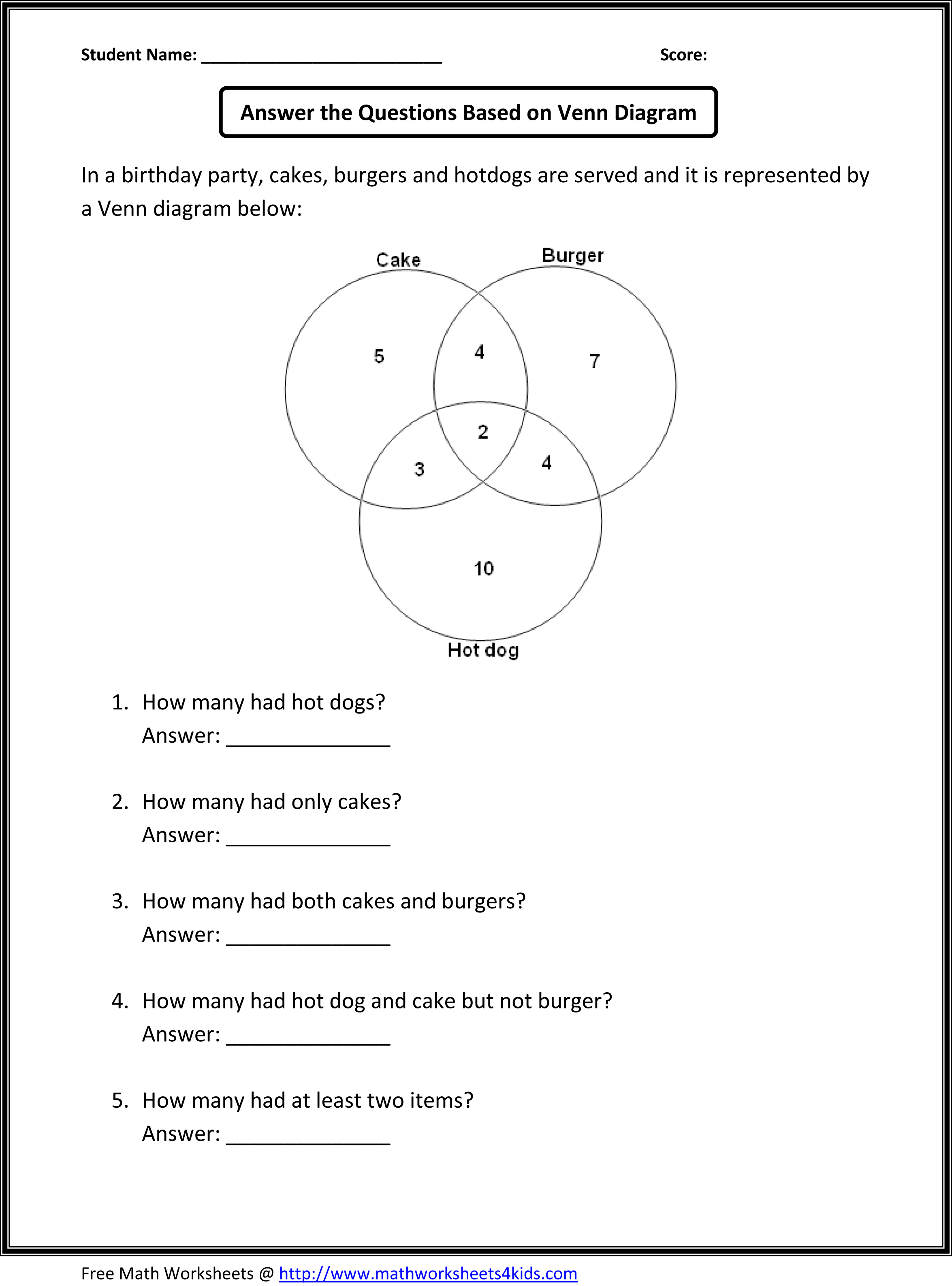  
One of the most popular methods to visualize the overlap and differences between data sets is the Venn diagram: a diagram consisting of two or more circles in which each circle corresponds to a data set, and the overlap between the circles corresponds to the overlap between the data sets. Often it is useful to see the overlap between different lists, enabling researchers to quickly observe similarities and differences between the data sets they are analyzing. In many genomics projects, numerous lists containing biological identifiers are produced.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
